Methods
Summary
The sampling for this project will take place at 3 National Estuarine Research Reserve System (NERRS) sites in Florida. I will be sampling at each site for 8 days in September of 2016.
I will be accompanying researchers, interns & volunteers at each site as they monitor sea turtle nesting beaches for post-hatch nests. At each beach, these nests are excavated, and the number of hatched vs. unhatched eggs are counted in order to assess hatchling success. As the monitoring groups excavate nests, will be taking a variety of samples:
1) Sterile swabs of unhatched eggs, eggshell fragments and dead hatchlings
While sampling, swabs will be kept in sterile microcentrifuge tubes. Once back in the lab, each swab will be plated onto a Fusarium - selective culture media called NASH agar.
NASH consists of:
· Distilled Water 500mL
· KH2PO4 0.5g
· MgSO4 0.25g
· Peptone 7.5g
· Agar 7.5g
· PCNB 0.5g
· Streptomycin sulfate 0.15g
· Neomycin Sulfate 0.06g
These plates will be sealed with parafilm and monitored for growth daily. Potential Fusarium isolates will be identified based on colony morphology, and aseptically transferred into pure culture for future morphological analysis.
2) Sand samples from within the nest and at set intervals outside of the nests
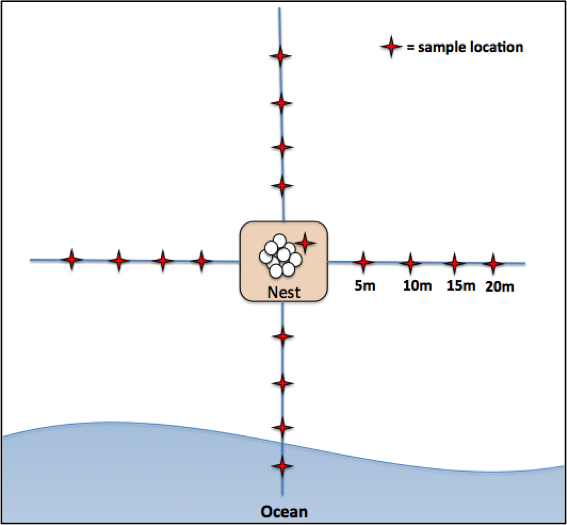
Sand samples will be collected from the inside of each excavated nest using a 10cm deep core and a spatula to transfer it to sterile whirlpack bags. Sand samples will also be collected from the surface, using a 10cm deep core, around each excavated nest at 5, 10, 15 and 20 meters in each direction parallel and perpendicular with the ocean (Figure 1). I will also collect sand from each nesting beach at least 50 meters away from any known sea turtle nests. This will allow for comparisons between nesting and non-nesting areas.
A dilution plating method (Bateman and Coskun 1995) will be used to assess Fusarium diversity, frequency and abundance from the sand samples. Single colonies of Fusarium will be sub-cultured into a pure culture for further morphological and molecular analyses.
The sampling design at each nest is representative of one unit in a larger hierarchical sampling scheme, encompassing multiple beaches within a NERRS site and multiple NERRS sites within Florida. Such a hierarchical design is ideal for preliminary determination of the genetic structure of populations of soil-borne fungi when very little information is known. This will ultimately help elucidate the movement of clones of Fusarium keratoplasticum and Fusarium falciforme in populations as well as any geographic boundaries of distinct populations (McDonald et al. 1997).
3) Ocean water samples at each beach
Surface ocean water will be collected at each nesting beach with sterile 50mL conical tubes. Water samples will be subjected to the same dilution plating method as sand in order to assess Fusarium diversity, frequency and abundance in ocean water.
Site Characteristics: At each sampled nest, data on the following characteristics will be collected: sand temperature inside and outside of nest, air temperature, soil composition, % vegetation, mean distance from water’s edge. Data collected on site characteristics will allow for comparisons to be made between nests sampled, as well as between sampling sites, with regards to the diversity, frequency and abundance Fusarium species.
All Fusarium isolates will be brought back to Penn State for further identification using molecular techniques. Preliminary molecular identification of Fusarium isolates will be accomplished using the preferred locus for Fusarium identification, a portion of the translation elongation factor 1-alpha gene (TEF; Geiser et al. 2004). All Fusarium keratoplasticum and Fusarium falciforme isolates will be analyzed using the 9-locus MLST developed by Short et al. (2014), to determine connections between nest-associated isolates and known isolates from anthropogenic environments, as well as other sea turtle nesting sites.
Challenges
Space for, and transport of, sampling equipment will most likely be my #1 challenge. I have chosen sampling devices and methods that should minimize the amount of space required to transport all of the necessary material. One way I plan to combat this issue is by shipping Fusarium cultures I have collected back up to Penn State for storage until I can return to the lab to process them.
I will also have limited space available at each site to complete my dilution plating, subculturing and microscope work. This will force me to be resourceful with the area I have while maintaining a sterile work space.
Another challenge I could face is weather. By law, excavation of sea turtle nests inundated by water during flooding events must be postponed for several weeks after the flooding event occurred. Any kind of major storm event could reduce the number of nests I am able to sample during my limited time in Florida. There is not much I can do about this other than cross my fingers & hope for the best!
Protocols
This project has not yet shared any protocols.