Please wait...
About This Project
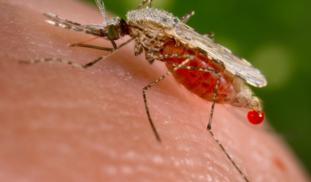
Yes, mosquitoes are flying syringes! They sample human blood as a normal part of their life cycle. We can take advantage of this nasty habit by using infected mosquitoes as a tool for monitoring the risk to human populations. Viral infection level in mosquitoes reflects the infection level in humans (Kilpatrick, 2013). Mosquitoes can also be used to screen for animal reservoirs of Zika. We’ve used this method to screen insects for parasites and are committed to adapting it to screen for Zika.
More Lab Notes From This Project

Browse Other Projects on Experiment
Related Projects
Disrupting cancer cell signaling through drug discovery
Most cancer-related deaths are caused by metastasis, the spread of cancer cells to distant tissues. This...
CaniSense– AI-powered blood test for early cancer detection in dogs
Cancer is the leading cause of death in dogs, yet no reliable methods for early screening exist. At testblu...
Shutting down cancer’s recycling system with exosome-based therapy
Pancreatic cancer is one of the deadliest cancers because its cells survive by recycling their own components...