About This Project
A major goal of ecology is to understand how organisms are distributed in nature. My research addresses the consequences of a low-dispersal red seaweed that inhabits a vast range of rocky shores across Western and Northeastern Atlantic coastlines. I will use genetic techniques to explain the relationship between short-distance dispersal and long-range distribution. This project will underscore the role of dispersal on the spatial scale of gene flow.
Ask the Scientists
Join The DiscussionWhat is the context of this research?
In marine habitats, water movement often drives spatial distribution of marine species through mediation of larval and spore dispersal (Selkoe et al. 2008). Studies examining the effects of oceanographic features on gene flow can distinguish genetic lineages across spatial scales (Gordiano et al. 2007). Intertidal seaweeds, for example, are often characterized by short dispersal distances due to a lack of longevity in dispersal stages and low levels of water movement (Kreuger-Hadfield et al. 2013). For the red seaweed Mastocarpus stellatus, short dispersal distances in tandem with a broad geographic range provide an excellent opportunity to investigate an interesting biological question: What is the spatial scale of genetic structure for a low-dispersal organism?
What is the significance of this project?
Most animals that occupy broad geographic ranges exhibit high levels of population connectivity due to life cycles involving long-term larval durations that allow movement of individuals between populations as far as 100 km in some cases. For broadly distributed populations of organisms with short-distance dispersal, however, the rate of increasing genetic divergence in relation to spatial distance is yet to be understood. Examining patterns of gene flow among populations at a massive range of geographic distances (10 cm – 1000 km) will provide an understanding of the spatial scale of genetic diversity in organisms that are seldom studied in research focusing on genetic relationships.
What are the goals of the project?
The overarching goal of my research is to identify the spatial scale of genetic diversity in the red seaweed Mastocarpus stellatus. To achieve this goal, I will be examining genetic diversity between populations at three relevant spatial scales: fine-, medium-, and large-scale geographic distances. I have collected fronds of this species from 11 sites along the coasts of Maine and France for extraction of genomic DNA. For each collected individual, exact spatial positions (± 1 cm) have been recorded, and GPS coordinates are available for each site. After completion of the final procedural step of collecting genetic data at 16 microsatellite loci for each frond, I will compare geographic distances with genetic distances.
Budget
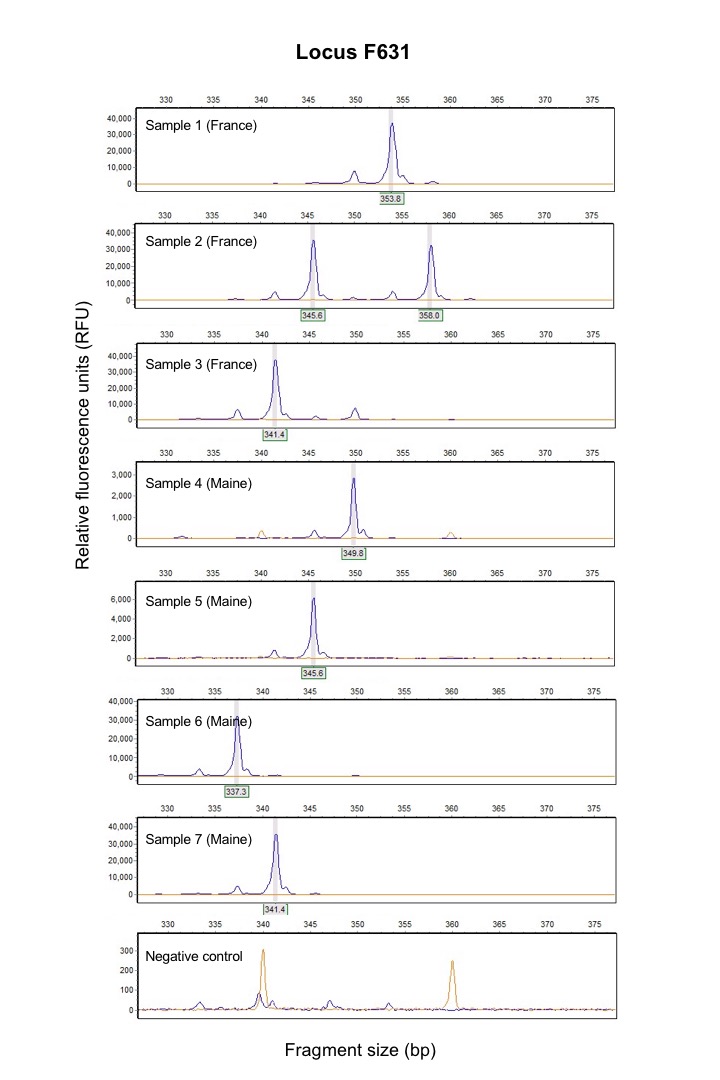
As of now, I have identified 16 genes that will allow me to estimate gene flow among populations of red seaweed. DNA has been extracted from the samples collected at each site from rocky shores along the coastlines Maine and France, and the remaining step is to isolate the 16 genes of interest for every sample (over 1,000 individuals). To achieve this goal, I will be using a technique called multiplex PCR, which allows me to develop short nucleotide sequences that target these genes in a controlled experimental procedure. I will add fluorescent tags to these sequences so that I can use a scanner to read the exact sizes of the genes that I am looking for. Each of the items in my budget contributes a small but indispensable role in completion of this part of the project that will result in genetic data that will then be interpreted through statistical analyses.
Endorsed by
 Project Timeline
Project Timeline
The funds for this part of the project will allow me to begin ordering the necessary supplies by the week of March 5. I will carry out multiplex PCR for every study site from March 5 – May 18. PCR products will be read using a scanner for fragment analyses from March 12 – May 31. I will then have the data collected, and will analyze the data from June 1 – August 31. Once the data is analyzed, I will write a manuscript that describes the project overall.
Feb 20, 2018
Project Launched
Mar 05, 2018
Order lab supplies
May 18, 2018
Complete multiplex PCR for every sample
May 31, 2018
Conduct fragment analyses for PCR products
Aug 31, 2018
Analyze the data
Meet the Team
Malek Al-Marayati
I have been working on this project as a Project Manager in the lab of Dr. Steve Dudgeon in the Department of Biology at California State University, Northridge for two years. I received my B.S. from CSUN in Marine Biology, and I hope to continue working on population ecology in the near future.
Additional Information
Project Backers
- 59Backers
- 106%Funded
- $4,932Total Donations
- $83.59Average Donation


