About This Project
Embryonic cells make choices that guide the differentiation of their clonal descendants. Such decisions establish the outline of a snail's whole body in a spatially patterned array of just twelve diverse, unambiguously identified single cells which are differentially exposed to a signal briefly emanating from a point source. We want to know if (and how) a change in this pattern causes the neck-founder cells to inappropriately form trunk structures, and if lack of a neck makes the head pop off.
Ask the Scientists
Join The DiscussionWhat is the context of this research?
How do embryonic cells decide to form various body regions, organs and tissues? Snail embryos build the body almost entirely from thirteen (easily recognizable and accessible) regional founder cells that are formed during the first few hours of development. Single-cell deletion experiments indicate that at these early stages, each cell becomes determined to form a certain body region. These decisions depend on cell-intrinsic factors in combination with an intercellular signal. In an unpublished experiment, removal of the right-hand body quadrant founder cell often leads surprisingly to development of a perfect trunk and a small, detached head. Why? We hypothesize that cell-intrinsic factors are acting normally in this situation, but the distribution of intercellular signal is altered.
What is the significance of this project?
Importantly, this study analyzes development by focusing on unique single cells with well-defined neighbor relationships and precisely predictable behaviors over time. We hypothesize that after early loss of the right-hand body quadrant founder cell, some remaining cells are abnormally exposed to an intercellular signal triggering ERK activation, and consequently adopt the right-hand quadrant fate. Another question: why does the head pop off? We hypothesize that cells inappropriately activating ERK (and forming trunk structures) are diverted from a 'neck' fate that normally plays a role in keeping the head attached. Defining the 'positional value' of each founder cell in an alternative situation, and connecting this with altered cell fates, will help define the elements of this system.
What are the goals of the project?
The basic experiment - ablation of one of the embryo's four quadrant-founder cells - was done a long time ago (Clement, 1956: J Exp Zool 132: 427-446), but the results were never reported. We have repeated this experiment and obtained reproducible results. Now we have three goals for further analysis. The first is clonal fate mapping, where a fluorescent dye injected into a single cell is inherited by descendant cells, showing what tissues and organs develop from the injected cell. Second, we want to analyze ERK activation at the single-cell level in early embryos, to test the idea that intercellular signaling is altered in three-quarter embryos. Third, we want to visualize molecular markers of early and later developmental pathways in the experimentally altered embryos.
Budget
A stereomicroscope with adequate (bright) light and (80x) magnification is essential for intracellular microinjection of dyes to map clonal fates of early embryonic cells. Hydrofluoric acid is needed to open the tips of microinjection needles. Antibodies are used for immunofluorescence visualization of ERK activation connected with normal cell fate specification. In situ hybridization is used to visualize RNA transcripts of genes activated in response to ERK signaling.
Endorsed by
 Project Timeline
Project Timeline
15 July 2020: Results published in poster (SDB 2020 meeting) - shared on Experiment.com
30 September: Troubleshooting, using HF to open needles, lineage tracer injection of 2b cell in whole and partial embryos.
30 October: New antibodies, try again to stain whole and partial embryos for active ERK.
30 November 2020: Data for ERK activation and clonal fate mapping hopefully piling up.
30 December 2020: Complete data sets for ERK activation and clonal fate mapping hopefully available.
Jul 14, 2020
Poster posted. Look forward to more comprehensive background and details of methods, results, and conclusions!
Oct 14, 2020
Project Launched
Oct 30, 2020
More data shared - ERK activation, lineage tracing...
Jan 30, 2021
Manuscript submitted for publication, hopefully!
Meet the Team
Morgan Q. Goulding
I am a developmental cell biologist and a teacher of wide-ranging topics in biology. I am especially interested in cellular processes underlying the earliest steps in patterning animal body plans.
My doctoral research applied single-cell deletion and other microsurgical interventions in embryos of the sea snail Ilyanassa, revealing that cell contacts NORMALLY shift the plane of a specific cell's asymmetric division, preventing its daughter cell from developing EXCESS neurosensory organs. In connection with this study, I also compiled a description of the Ilyanassa embryo's spatiotemporal pattern of cell division up to the 84-cell stage, revealing in this species a widespread tendency toward increased complexity compared with analogous patterns in other kinds of snail embryo.
As a postdoctoral researcher, I extended my explorations of cellular mechanics in early development by using genetics and live cell imaging analysis to dissect the spatial regulation of forces that move centrosomes in the C. elegans zygote.
More recently I have continued exploring how early embryonic cell cycles are regulated in coordination with the earliest cellular decisions impacting developmental potential in gastropods, which offer unique advantages as a model system for this area of discovery.
Additional Information
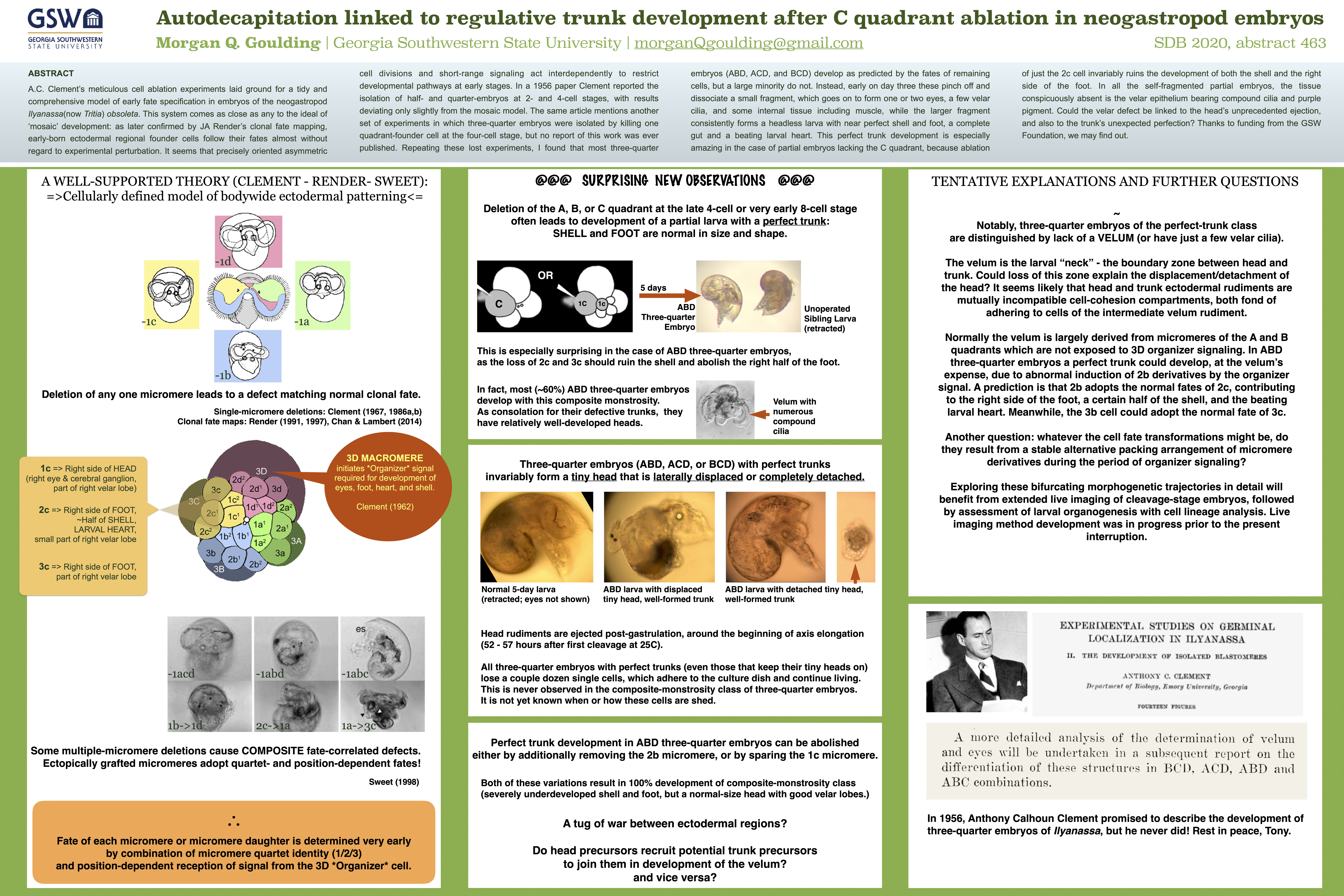
A poster documenting the early steps in this project was presented at the 79th Annual Meeting of the Society for Developmental Biology, July 2020.
Project Backers
- 12Backers
- 100%Funded
- $1,510Total Donations
- $125.83Average Donation

